Dragon
Dragon. Spray + Acrylic on canvas. Stencils built using a CNC shopbot.
Dragon. Spray + Acrylic on canvas. Stencils built using a CNC shopbot.
When building crawlers, most of the effort is expended in guiding them through a website. For example, if we want to crawl all pages and individual posts on this blog, we extract links like so:
In this blog post, I present a new DSL that allows you to concisely describe this process.
This DSL is now part of this crawler: https://github.com/shriphani/pegasus

The incredibly talented Pem Lasota, Aram Ebtekar and I put together a small project to enhance the music experience in our living room.
What ensued is the first of many projects we plan to roll out - all focused on building the greatest living-room music experience in the world.
The setup involves 4096 LEDs arranged in a matrix, powered by a raspberry pi and a beefy condenser mic. The electronics fit into a 3D printed case. Pem did a significant chunk of this in Solidworks - a piece of software I was extremely impressed with. In one of the sessions, I was able to make reasonable headway with mild supervision. Few pieces of software are this easy to pick up. Solidworks have done a solid job.
and when everything is put together, it looks like this:
The frame listens via the condenser mic and generates beautiful visualizations when it “hears” audio. This is what happens when we blast Pavarotti on our music system:
A neutrino effect to Flume's Hyperparadise:
The Matrix effect when the bass drops:
In India, 30% of goods sold by foreign companies must be manufactured within the country.
This law puts a damper on Apple’s India plans - threatening to prevent them from bringing their incredibly successful retail strategy to the country.
Turns out there’s an exemption for retailers providing cutting-edge technology. Apple apparently failed to qualify for this exemption. This decision is top-level comedy coming from a country that has failed to deliver indoor plumbing to more than half its citizens.
There’s a beautiful anecdote from the 80s or 90s (or some other time the nation was conducting its large-scale economics-voodoo social experiments). Sunil Mittal - founder of Airtel, one of India’s largest telecom companies - saw a push-button phone on a trip in Taiwan and decided to bring it to India - where the rotary dial was still state-of-the-art.
Turns our phone imports were banned. The burgeoning company had to set up operations to buy fully built phones in Taiwan, break them apart, ship them to India, and then reassemble them.
I am sure that this lunacy was fully paid for by the customers.
Sunil himself mentions this story near the 16:25 mark:
If you wondered where to look for a Yakov Smirnoff jokes in a post Soviet world, India’s got your back.
India’s vast bureaucracy is now bringing its finely honed judgement to deal with the world’s most successful companies.
core.cache is a small and convenient cache library for clojure. It enables clojure users to quickly roll out caches. In this blog post I am going to describe a clojure implementation which stores cache entries to disk: fort-knox.
In a few recent projects I’ve needed a cache with entries backed to disk. This is a vital requirement in applications that need to be fault-tolerant. LMDB (which I’ve had very positive experiences with) is fast, quick and perfect for this task. fort-knox implements the core.cache spec and stores entries in LMDB. clj-lmdb (subject of a previous blog post) is part of the plan now.
Note that this library deviates slightly from suggestions for core.cache implementations. For instance, the backing store doesn’t implement IPersistentCollection or Associative so fort-knox might deviate from expected behavior. Thus YMMV.
Quartz has a new news app - an absolute peach.
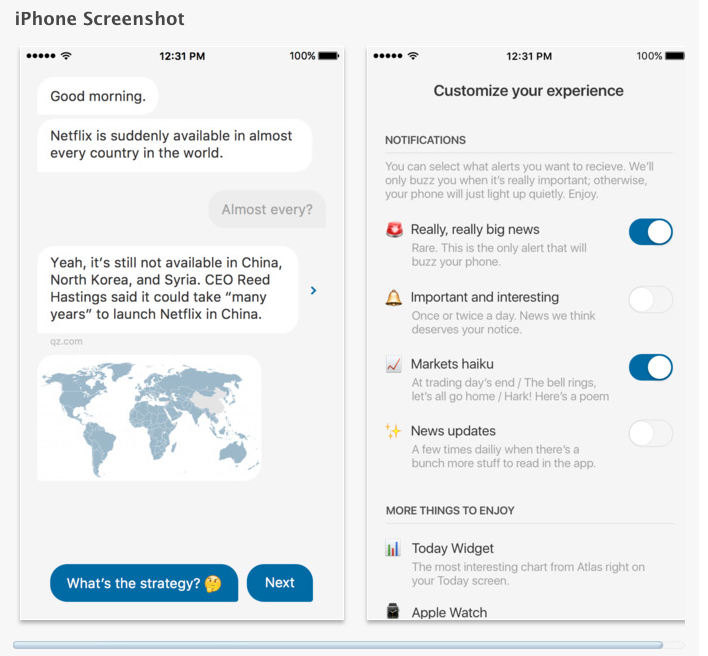
The NYTimes in comparison is a brain-dump.

As a kid, in the pre-WWW days, people around me consumed news in the following forms (I couldn’t be bothered):
The WWW presented a challenge (and still does) for traditional news orgs. I believe all content gets 2 kinds of eyeballs - the kind of folk who are looking for exactly that content and the kind who are bored and will watch whatever is on. When a TV producer sets up a narrative for a day’s worth of programming, you get the attention of both these groups - there are few other options.
In an on-demand world, long-form news that is hastily put together needs to compete with youtube videos, reddit and so on. Converting a narrative of events into a blob of text you can put on your site is going to attract only the kind of people who want to read this sort of stuff in their spare time.
In my experience this species loves the color beige and has the world-view of prep-school headmasters.
And to capture and lock down this audience, you are forced to convert journalism into a weird vaudeville performance. In fact The Daily Show ended up becoming a meta-news org making fun of this phenomenon.
As print journalism and 24 hour news orgs die, I see 2 main vehicles for news consumption emerging:
This leaves us with how to make $$.
I don’t think breaking news is a serious vehicle to make money (anymore). However existing media shops need to invest in this space to maintain an aura of legitimacy. This might change in the future but today’s expectations of a news org include breaking news. It might be ok for a newer media org to become purely an analysis-based-reporting organization (like Nate Silver’s http://fivethirtyeight.com/), but I would still like the news app to alert me when something serious happens (like the Paris attacks) and not have to discover this on Twitter.
Quartz has been delivering small ads in the news stream that look like this:

I like these sorts of ads - they aren’t intrusive (well, for now) and possibly get you more impressions than whatever the status quo is.
There are 2 additional examples I want to cover. TomoNews is an incredibly creative news org. The animated shorts are rather entertaining.
BuzzFeed has mastered the art of bringing people to their site, creating a stable revenue stream from their baity content and produce some very nice long-form news articles.
To survive, journalism needs to change. There are new gatekeepers, new platforms and new competitors. News needs to change from a brain-dump to a new kind of performance art. Existing media powerhouses have several weapons in their war chest - a brand-name that resonates well, existing relationships with the rich and powerful, and revenue streams (they might be dwindling but at least they’re around).
This soft-power needs to mix with the best photography, web-design, typography and visualization-tech. Each piece of investigative journalism should become a distinct, pleasant memory - like when Walter Cronkite covered the moon landings:
In short, give people something they will enjoy for a reasonable price.
In 2014 I attended a talk by Ramanathan V. Guha on schema.org. Its remarkable adoption offers several lessons about building internet properties.
Schema.org is an ontology - a vocabulary for annotating HTML documents with information. For instance, you can put in some info in the web-page markup stating that what you’re displaying is blog post or a list of products.
For instance, a website can present its content and state that this content is an instance of Blog (say). Schema.org is a central repository describing what Blog is, what and where it fits in an ontology.
As of 2015, Schema.org is by far the most dominant vocabulary used on the internet [1].
Schema.org accomplished this feat by aligning incentives correctly. First, annotated webpages would result in a richer search-result presentation - translating to better traffic to the website - a massive economic incentive. Next, the top 4 players in search agreed to support Schema.org - Google was obviously the mothership, bing, Yahoo and Yandex supported it.
Schema.org offers an amazing lesson in setting up the incentives correctly. When you’re building a standard, waiting for people to adopt your rules on the basis of their merit is slightly less efficient than not building the standard at all. A clear economic incentive is a powerful motivator.
The Schema.org team understood that well. And now there is an amazing foundation that is being exploited by a new generation of applications.
LMDB is the nicest, no-nonsense, no-surprise key-value store I’ve ever used.
In several benchmarks, LMDB destroys competitors - it is a beloved tool in high-profile circles.
Here’s clojure bindings: https://github.com/shriphani/clj-lmdb
One of the more memorable courses I took at Carnegie Mellon was Manuel Blum’s algorithms.
The CSD made it a mandatory course for some major (I don’t know about this since I was just fucking around for the most part).
It is a little known secret that you don’t attend Manuel’s course to learn about algorithms - you can learn quite well by opening a book.
You attend to watch a genius combine expertise, brilliance, and passion.
When I took the course, Manuel wanted to get up to speed with machine learning. As of 2014 Manuel had won the Turing award, his students shared among them three Turing awards, founded new fields in computer science like Quantum Computing, and founded successful companies which went on to become cultural phenomena - ReCaptcha and Duolingo among them.
Manuel’s course essentially involved delivering a lecture on a topic of your choice - I made a proper horlicks of my talk. However, it inspired a lively blog post and visualization here.
I was pretty excited to meet Manuel in person. Back in high school I was avid follower of Scott Aaronson’s blog and he spoke highly of his advisor Umesh Vazirani. Manuel was Umesh’s advisor and it was a meeting-your-childhood-hero moment for me.
From the entire course, two things stood out:
Manuel has tons of amazing advice for students on his website. Personally, when I took his course, due to a variety of circumstances, I had mentally checked out of CMU.
Manuel was the display of intellect I needed to not descend into deep cynicism about a life devoted to learning.
#KeepPounding #Panthers #SB50 Super Bowl 50: Broncos vs. Panthers CBS https://t.co/BwlJmzjPlX pic.twitter.com/iNWUm6lgWs
— ClippitUsers Sports (@FanSportsClips) February 8, 2016
Twitter: @shriphani
Instagram: @life_of_ess
Fortior Per Mentem
(c) Shriphani Palakodety 2013-2020